The authors separated the metastatic breast tumour cells into active and resting phenotypes depending on CD90 positivity, with high or low scatter respectively. When co injected together with the adipose derived MSCs into mice, only the lively effusion cells were tumourigenic. Park and co employees reported the migration of human umbilical cord MSCs in direction of human glioma cells in vitro, and that overexpression of CXCR4 elevated this trait. Even more, in a xenograft model of glioma in nude mice, these cells displayed enhanced migration to the tumours. In an experiment in mice using transplan tation of GFP tagged BM, GFP good MSCs migrated into the prostate of castrated mice, and these cells were greater by testosterone in a Wnt dependent manner.
These findings had been also noticed within a human prostate tumour xenograft, during which MSCs expressing an exo genous Wnt antagonist, secreted Frizzled connected protein two, induced tumour shrinkage by necrosis and apoptosis. Kucerova and colleagues reported that adipose tissue MSCs could promote growth additional reading in nude mice of tumours on the xenografted human melanoma cell line A375. This was accomplished by suppression of apoptosis and a rise in proliferation. An additional melanoma line, 8MGBA, didn’t share this attribute, rather, MSCs have been inhibitory. Two recent reports propose that MSCs may perhaps give rise right to mesenchymal tumours. Making use of a comparison of infused regular MSCs, in vitro spontaneously trans formed MSCs, and osteosarcoma murine cells, Mohseny and co workers concluded that aneuploidy, chro mosomal translocations and also the homozygous loss of the Cdkn2A locus on chromosome four were implicated in tumour progression.
The genetic alterations seemed to come about about MSC passages 5 to 9 in culture, throughout which time the cells went into crisis, and thereafter they possessed the capability to expand in soft agar indepen dently of substrate. The authors showed a series of 88 human osteosarcomas that possessed comparable defects from the selleckchem homologous cyclin dependent kinase inhibitor 2A locus on chromosome 9. Kaplan Meier ana lyses of these individuals with osteosarcoma showed extremely bad survival for individuals unfavorable for this locus. Though this examine did formally show the origins of these human osteosar comas to be MSCs, it warrants a cautious technique when making use of these cells from the clinic. A further report of tumours arising from genetically defective MSCs has lately appeared.
These authors deleted p53, Rb or the two genes in adipose tissue derived murine MSCs that underwent Cre LoxP exci sion. Wild kind and Rb negative MSCs were phenotypi cally standard, whereas the p53 unfavorable and p53 negative/Rb unfavorable MSCs were transformed, 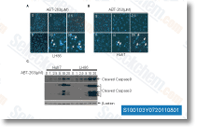 and could initiate leiomyosarcomas in half the animals when injected in to the flanks of NOD SCID/IL 2Rg mice. The transformed MSCs approached tetraploidy, and had been deficient during the capacity to differentiate into adipo cytes, still had greater means to become osteocytic.
and could initiate leiomyosarcomas in half the animals when injected in to the flanks of NOD SCID/IL 2Rg mice. The transformed MSCs approached tetraploidy, and had been deficient during the capacity to differentiate into adipo cytes, still had greater means to become osteocytic.
Monthly Archives: May 2014
This structure proficiently creates an additional amount of robus
This structure efficiently generates an extra level of robustness of cell fate dedication, and that is rendered by two new styles of bistable switches, additionally towards the reprogramming switch. 1 kind of switch includes the 2 bistable areas positioned at lower variety on the primary signal, which controls differentiation/dedifferentiation commitment, i. e. the switches from or for the na ve state. Yet another type of switch consists of the 2 bistable areas situated at larger array in the principal signal, which controls co expression commitment, i. e. the switches from or on the double positive state. We define these two switches because the differentiation switch and the co expression switch respectively.
The tri steady regions within this diagram are the overlapping regions among the selelck kinase inhibitor bi steady areas governed from the reprogramming switch and both the differentiation or the co expression switch. In reality, really large weights of car activation might give rise to a tetra stable area, the place the three sorts of your bistable regions overlap. In summary, the favourable feedback loop involving mu tual inhibition in the master regulators can develop the re programming switch, and further feedback loops involving auto activation can improve the robustness in the reprogramming switch and make the differentiation switch and also the co expression switch. The attributes on the three bistable switches are listed in Table 3. We subsequent ran simulations to check out whether or not these areas of multistability are correlated to many sorts of heterogeneous differentiation.
Our outcomes present that Sort 1 heterogeneous differentiation might be induced from the reprogramming switch region, Sort 2 heterogeneous differentiation may be induced from the co expression bistable switch areas, and Type three heterogeneous differentiation can be induced inhibitor PI-103 while in the tri stable region consisting of 3 func tional states. These kinds of het erogeneous differentiations are all robust when it comes to single cell commitment due to the fact the corresponding par ameter areas admit a variety of secure regular states. Positive suggestions loops have long been acknowledged as mechanisms for biological switches. We’ve got demonstrated that two styles of favourable feedback while in the CD4 T cell differentiation network underlie three forms of bistable switches that govern the transitions among unique phenotypes of people T cells.
Additionally to en suring the robust commitment, the multistability created by beneficial suggestions loops can be applied to make phenotypic diversities of different varieties. Within this context, the biological functions of the constructive feedback loops are witnessed as additional versatile than offering rise to basic on or off switches. Our theoretical analysis on the basal regulatory motif began with symmetrical parameter values then viewed as the results of broken symmetries.
However, a study finished by Park et al suggests implication of
On the other hand, a review done by Park et al. suggests implication of PLD in the signaling cascade that activates COX two, leading to professional duction of prostaglandins and initiation of labor. Considering that all ureaplasma serovars and also the four sequenced clin ical isolates consist of a gene with PLD domains, a long term functional characterization of this gene would be of inter est. We’ve not been in a position to find computationally the genes encoding PLA1 and PLA2 in ureaplasmas. IgA Protease In the mammalian immune procedure, a principal defense mechanism at mucosal surfaces is the secretion of im munoglobulin A antibodies. Destruction of IgA anti bodies by IgA distinct protease lets evasion of the host defense mechanism. In Neisseria gonorrhoeae the IgA professional tease doubles as a LAMP one protease to permit it to stop fusion of the phagosome with the lysosome. IgA professional tease action was demonstrated in ureaplasma serovars.
All sequenced human ureaplasma genomes were evaluated for IgA protease genes with all the same approaches since the phospholipases selleck inhibitor gene search.We couldn’t compu tationally recognize an IgA protease gene. Nucleases Nucleases are actually reported as possible pathogenicity variables in other organisms at the same time. Ureaplasmas be prolonged to a group of organisms that import nucleotides for DNA and RNA synthesis. Therefore it is actually very likely they have secreted or surface bound nucleases that may also perform a purpose in pathogenicity. We recognized 15 likely nucleases, of which two had a predicted signal peptide, and therefore are prone to be secreted or surface bound. These nucleases could be an fascinating target for more studies of their potential involvement in pathogenicity. Putative O sialoglycoprotein peptidase Eleven from the 14 ureaplasma serovars contained a gene annotated as an O sialoglycoprotein endopeptidase.
UUR serovars two, 8, and ten did not include an ortholog of this gene. Since all three of those genomes are complete, we will be certain the gene is absent. This enzyme has been shown to cleave human erythrocyte glycophorin A in other bacteria. Precisely the same study showed that the specificity of this peptidase is restricted to O glycosylated membrane glycoproteins, and it are not able to cleave N glycosylated buy R547 proteins. Abdullah et al. suggest that the possible targets of this enzyme during the host are sialoglycoproteins in the mucosal epithelial cells or over the cell surfaces of macrophages. In actual fact the O sialoglycoprotein peptidase of Mannheimia haemolytica cleaves in the surface of the human cell line KGla the CD43 leukosialin along with other human O sialoprotein anti gens like the progenitor cell restricted antigen CD34, the hyaluronate receptor CD44, plus the leukocyte typical antigen tyrosine phosphatase CD45 class of molecules.
The pteridine pathway is uncovered in both plants and animals alo
The pteridine pathway is identified in both plants and animals and a key compound from the pathway, tet rahydrobiopterin, acts as an critical cofactor during the degradation of phenylalanine and also the synthesis in the neurotransmitters serotonin, melatonin, dopamine, nor epinephrine and epinephrine, Pteridine and ommo chrome pigments type the basis from the noticeable eye color variants of Drosophila and a great deal on the variation in butter fly wing patterns, and have consequently been central towards the advancement of genetics itself, Indeed the plethora of observed eye color mutants in Drosophila benefits through the complex spectral interactions of pteridine and ommo chrome pigments. Offered the use of guanine being a colorant in spiders, it really is also exciting to note that this is often the key substrate for the pteridine pathway, Lastly, many pigment proteins contain heme groups or end result from conjugates of heme containing compounds, The parallel evolution of genetically based adaptive modifications amongst both unrelated species along with the hugely structured populations of these spiders tends to make these programs great for examining evolution beneath balancing assortment.
Our ultimate aim is usually to elucidate the molecular basis of your evolutionary alterations which have led towards the parallel evolution of comparable coloration in these species. On the other hand, a important stage on this system is the determination of your pigment synthesis pathways which might be existing in these spiders plus the gene sequences connected with them. Subsequently candidate genes selleck chemicals linked with the allelic basis in the colour polymorphism or that are differentially expressed amongst color morphs might be recognized.
The advent of following generation sequencing technologies has permitted speedy profiling and de novo assembly of the total set of expressed mRNA se quences in a unique tissue or complete organism, Moreover to giving data about the framework of expressed gene transcripts, the digital nature of RNA seq facilitates the determination of the two relative order DMXAA transcript expression ranges inside of a tis sue or organism and also the differential expression of tran scripts among tissues or experimental therapies. Applying data created through a blend of RNA seq and the sequencing of normalized cDNA libraries to com pensate for your underneath sampling and poor assembly of rarer transcripts, we report to the close to finish total body expressed transcriptomes of two species of shade polymorphic spider, Theridion californicum and T. grallator. This represents quite possibly the most extensive genomic information set for spiders so far offered. We report to the gene complement of those species and highlight gene families that appear to possess seasoned expansion in the lineage resulting in spiders.
glabripennis has the metabolic probable to overcome numerous of t
glabripennis has the metabolic possible to conquer quite a few from the problems linked with feeding in woody tissue, such as degrad ation of lignin, cellulose, and hemicellulose and acquisition of nitrogen and various important nutrients, the contribu tions of insect derived digestive and nutrient obtaining enzymes can’t be ignored given that insects themselves can create a diverse array of digestive enzymes, including cellulases, hemicellulases, pectinases, and enzymes that boost lignin degradation, Insects have also evolved other sophisticated skills to evade host plant defenses and generally possess in depth suites of enzymes concerned in detoxification of plant metabolites and phyto hormones, digestive proteinase inhibitors, and cyanates and cyanoamino acids, likewise as enzymes capable of disrupting jasmonic acid signaling pathways, Even more extra, insects produce numerous cytochrome P450s, that are integrally concerned in xenobiotic metabolic processes that in the long run lead to oxidative destruction of toxic com pounds, like plant derived secondary metabolites and pesticides, The main objectives of this examine were to survey the endogenous digestive and physiological abilities of larval A.
glabripennis by shotgun sequencing of midgut derived messenger RNA and to determine insect derived genes which might be very expressed while in the midgut whilst actively feeding in wood. The A. glabripennis midgut transcriptome library was also compared inhibitor price to all publically accessible transcriptome libraries sampled from other plant feeding insects to recognize core groups of genes that are associated with digestive processes that may facilitate nutrient recovery from woody tissue regardless of insect taxa.
This examine represents an import ant addendum towards the rising database of genomic and transcriptomic resources available for coleopterans and fills a vital gap, representing the very first transcriptome sampled from a wood feeding cerambycid as well as the initially extensive evaluation of endogenous genes connected with wood feeding in insects. read what he said These findings give distinctive possibilities to bioprospect for enzymes that may be exploited for cellulosic biofuel production or other indus trial processes, and also to develop novel management approaches for this destructive wood boring pest and also other wood feeding insects. Outcomes and discussion 454 and Illumina Based Transcriptome Sequencing To produce a complete profile on the endogenous digestive and physiological capabilities of the. glabripennis, mRNA was collected from your midguts of third instar larvae feeding in the heartwood of the preferred host and was sequenced applying the two Roche 454 pyrosequencing and Illumina technologies.
Once more, R2R3 MYB transcription component MYB12 in the thalian
Yet again, R2R3 MYB transcription issue MYB12 within a. thaliana has become shown to perform like a flavonol certain activator of fla vonoid biosynthesis, Transcriptional regulation of fla vonoid biosynthesis, a major branch of phenylpropanoid pathway, managed by a set of R2R3 MYB transcription elements, have already been reported in various plants such as Prunus persica, Epimedium sagittatum at the same time, Apart from this TF, 18 transcripts coded for bHLH TFs have already been recognized right here. The bHLH domain with the maize R gene is reported to take part in anthocyanin formation and serve being a website link in between flavonoid formation and his tone modification, Amongst the varied functions, bHLH transcription elements also regulate the biosynthetic pathway of flavonoids, in a number of plant species, 1 DOF loved ones TF is recognized in our analyses.
AtDOF4 is recognized to influence metabolism in an environmental and tissue certain method by positively regulating the manufacturing of hydroxycinnamic acids in the hypocotyl and flower buds, and negatively regulating fla vonoid biosynthesis in pollen grains, With each other, TFs recognized here and connected for the phenylpropanoid path way may be explored even further Thiazovivin solubility during the regulation of podophyl lotoxin biosynthesis. In silico SSR marker identification SSRs can be divided into genomic SSRs and EST SSRs. EST SSRs are extra evolutionary conserved than non coding sequences and therefore possess a reasonably substantial transferability, Subsequent generation sequencing has recognized EST SSRs in lots of plant species, Having said that, there are actually no reports of EST SSRs in P. hexandrum to date.
SSRs have been identified CP-690550 price with MISA search instrument, which can be according to the criteria that a dinucleotide or maybe a trinucleotide pattern should really appear not less than 6 times, and tetra, penta and hexa nucleotide patterns must seem five instances each and every, SSR distribution and SSR mining of transcripts identified a total of 1,011 SSRs from 40,380 transcripts, with 94 transcripts containing a lot more than one particular SSR. By far the most abundant repeat form was dinucleotides and also the dominant tandem repeat motifs have been 6 and seven representing 19. 4% and 25. 7% respectively. Transcriptome wide survey of miRNA targets in P. hexandrum cell cultures MiRNAs are recognized to manage many developmental and effector genes on the posttranscriptional level, Utilizing oligonucleotide arrays, miRNAs are already proven to be differentially expressed amongst tissues and during the maturation in the fruit during the grapevine, Wong et al, predicted 3 wood connected genes, flavonol synthase like, xyloglucan fucosyltransferase and glucan synthase like genes to become the targets of miR170, miR172 and miR319, respectively, and suggested that these miR NAs could be immediately concerned in regulation on the phe nylpropanoid pathway and hemicellulose biosynthesis pathway.
Also, pathways through the key me tabolism, just like the citric
Also, pathways in the principal me tabolism, which include the citric acid cycle are down regulated throughout leaf miner defense response in resistant plants, a profile observed also in conifers, Outcomes of this research deliver evidence that most genes encoding enzymes in the citric acid cycle are down regulated in resistant plants, In the parallel ana lysis, a metabolite profile was established for resistant and susceptible genotypes employing an NMR primarily based approach and indicated that malate levels on resistant leaves are decrease than in vulnerable ones. Malate success from conversion of either fumarate or glyoxylate. Expression of fumarase, that converts fumarate into malate, is down regulated at T0 in resistant genotypes, what could describe the minimal malate amounts.
Production of malate from glyoxylate may additionally be deficient in resistant plants, when genes encoding for malate synthase, that converts glyoxylate into malate, and read full article for isocitrate liase, the upstream enzyme that converts isocitrate into glyoxylate and succin ate, are the two down regulated, In contrast to this profile, vulnerable plants exhibit a ordinary expression amounts for these genes at T0. As a result, the two metabolic and transcriptional profiles assistance the affirma tions that citrate cycle is down regulated in leaf miner resistant coffee plants, and the model of down regulation of primary metabolic process in herbivore resistant plants, Biosynthesis of JA starts with alpha linoleic acid release in non injured tissues, triggered by systemin and phospho lipase A2.
Alpha linoleic acid is then converted to JA after enzymatic procedures performed by 13 lipoxygenase, allene oxide synthase, jasmonate o methyltrans ferase and other people, A few genes through the JA biosynthesis pathway are up regulated in resistant plants at T0, together with these from downstream kinase inhibitor amn-107 steps for example jasmonate o methyltransferase which expression is ten fold increased than in vulnerable plants. All genes within the JA biosynthesis are both down regulated or up regulated at later on stages in susceptible plants, as as an illustration expres sion of LOX increases only at T1. These observations propose that the JA signalling pathway, including intermediate signaling molecules for instance oxo pentenyl cyclopentane, could possibly be impaired in suscep tible plants. Down regulation of genes from later techniques of JA biosynthesis, for example allene oxide cyclase, allene oxide synthase, carboxyl methyltransferase, the enzyme that converts jasmonic acid into methyl jasmonate, is observed at T1 and T2 in resistant plants. Hence, a suggestions regulation may be activated, using a re programming of transcriptional response on lea miner infection. f
Ohno Machado may be the Associate Dean for Informatics and Techno
Ohno Machado certainly is the Associate Dean for Informatics and Technologies in the School of Medication, University of California, San Diego, the founding Chief from the Division of Biomedical Informatics, and a Professor of Medication. She is an elected fellow in the American Institute for Medical and Biological Engineering, the American College of Health care Informatics, plus the American Society for Clinical Investigation. She may be the Editor In Chief on the Journal with the American Medical Infor matics Association. Her investigation focuses on predictive modeling, particularly which include the evaluation of indi vidualized probabilistic estimates for risk assessment and prognosis. Genetic variation modulating drug response. discovery and implementation Dr.
Roden introduced BioVU, a resource that back links URB597 clinical trial DNA extracted from clinically obtained blood samples to their de recognized electronic medical record, BioVU not only allows the dis covery of new genomic variants related with distinct clinical phenotypes, but additionally new phenotypes connected with unique genotypes in an approach Dr. Roden and his team termed phenome broad association examine, Dr. Roden also presented the Vanderbilt PREDICT system that empowers patients and health professionals using the genetic infor mation wanted to predict and aid avoid adverse unwanted effects of medicines. Dr. Roden served since the Director in the Vanderbilt Arrhythmia Service, the director from the Division of Clinical Pharmacology, and in 2006 was named the Assistant Vice Chancellor for Perso nalized Medicine. Dr.
Roden continues to be elected to member ship in the American Society for Clinical Investigation as well as Association of American Physicians, and he’s a fellow from the American Association Bicalutamide clinical trial for that Advancement of Science. An intrinsically disordered protein Swiss Knife like toolkit for signaling diversification Dr. A. Keith Dunker provided a complete assessment of intrinsically disor dered proteins and their critical position as a multifa ceted, Swiss Knife like toolkit that permits swift diversification of cell signaling to facilitate the improvement of metazoans and their fast evolution. Dr. Dunker is really a Professor of Biochemistry and Molecular Biology at Indiana University, in which he launched the Center for Computational Biology and Bioinformatics and served as its Director. He is very best identified for his investigation in understanding IDPs making use of bioinformatics approaches and laboratory experiments. He and his collaborators were the 1st to think about these professional teins as a distinct class with crucial biological functions. Genome sequences of wild and domestic bactrian camels Dr. Yixue Li presented draft genome sequences from the two a wild along with a domestic Bactrian camel.
cerevisiae Cet1 The yeast triphosphatase includes a nov el terti
cerevisiae Cet1. The yeast triphosphatase has a nov el tertiary framework by which the lively web page is located inside a topologically closed hydrophilic tunnel com posed of eight antiparallel strands, that are conserved in CaCet1 and Pct1, Mutational evaluation of Cet1 has recognized 15 personal side chains inside the tunnel which can be essential selleck chemical PP242 for Cet1 function in vitro and in vivo. Every single of your 8 strands contributes no less than one practical group to the energetic web-site. Mutational evaluation in the Cand ida triphosphatase recommended strongly the tunnel fold along with the constituents of the lively web page are similar, if not identical, in Cet1 and CaCet1, Here we address the essential query of irrespective of whether RNA triphosphatase is essential for cell growth in fungal spe cies aside from S. cerevisiae.
This really is not a straw guy is sue, given that S. cerevisiae encodes two homologous RNA triphosphatases, of which only Cet1 is original site necessary for capping and cell viability, We use classical genetic approaches to demonstrate that the respective genes encoding RNA triphosphatase and RNA guanylyl transferase are crucial in S. pombe. Utilizing a novel process of Enloe et al. to check gene perform in diploid C. albicans, we have been not able to disrupt each copies in the CaCET1 gene, signifying that RNA triphosphatase is additionally essential in that species, a substantial human pathogen. Primarily based on these findings, plus the presence of the Cet1 ho molog from the Apergillus fumigatus proteome, we con clude that RNA triphosphatase can be a valid target for antifungal drug advancement. Outcomes RNA Triphosphatase and RNA Guanylyltransferase are Es sential in S.
pombe S. pombe RNA triphosphatase Pct1 is usually a 303 amino acid polypeptide which has a homodimeric quaternary construction, The pct1 gene contains just one intron inside of the open reading through frame, S. pombe RNA guanylyltrans ferase Pce1 is a 402 amino acid monomeric protein, there aren’t any introns inside of the pce1 gene. Despite the fact that re combinant  Pct1 and Pce1 enzymes happen to be purified and characterized biochemically, and proven to function in cap formation when expressed in S. cerevi siae, there are already no antecedent genetic scientific studies in the essentiality of Pct1 or Pce1 in fission yeast. Right here we constructed pct1 and pce1 plasmids have ing five and 3 flanking genomic sequences through which the en tire triphosphatase or guanylyltransferase coding sequence was deleted and replaced through the kanamycin re sistance gene, The pct1.kanMX and pce1.kanMX constructs had been transformed separately right into a diploid strain of S. pombe and chromosomal integrants include ing one particular copy from the wild sort gene and certainly one of pct1.k
Pct1 and Pce1 enzymes happen to be purified and characterized biochemically, and proven to function in cap formation when expressed in S. cerevi siae, there are already no antecedent genetic scientific studies in the essentiality of Pct1 or Pce1 in fission yeast. Right here we constructed pct1 and pce1 plasmids have ing five and 3 flanking genomic sequences through which the en tire triphosphatase or guanylyltransferase coding sequence was deleted and replaced through the kanamycin re sistance gene, The pct1.kanMX and pce1.kanMX constructs had been transformed separately right into a diploid strain of S. pombe and chromosomal integrants include ing one particular copy from the wild sort gene and certainly one of pct1.k
The Trinity de novo RNAseq as sembly pipeline was executed using
The Trinity de novo RNAseq as sembly pipeline was executed applying default parameters, implementing the Lower flag in Butterfly and making use of the Jellyfish k mer counting method, Assembly was completed in three hrs and 13 minutes on a compute node with 32 Xeon 3. one GHz cpus and 256 GB of RAM on the USDA ARS Pacific Basin Agricultural Analysis Center Moana compute cluster, Assembly filtering and gene prediction The output on the Trinity pipeline can be a FASTA formatted file containing sequences defined like a set of transcripts, which include alternatively spliced isoforms determined for the duration of graph reconstruction while in the Butterfly phase. These tran scripts are grouped into gene parts which repre sent many isoforms across a single unigene model.
When many complete length transcripts have been anticipated to become present, it’s most likely that the assembly also consisted of er roneous contigs, partial transcript fragments, and non coding RNA molecules. This collection of sequences was thus filtered to recognize contigs containing full or close to total length transcripts or possible coding regions and investigate this site isoforms which are represented at a minimum level primarily based off of read through abundance. Pooled non normalized reads have been aligned for the unfiltered Trinity. fasta transcript file using bowtie 0. 12. seven, through the alignReads. pl script distributed with Trinity. Abundance of each transcript was calculated making use of RSEM one. 2. 0, utilizing the Trinity wrapper run RSEM. pl. Through this wrapper, RSEM read through abundance values had been calculated on the per isoform and per unigene basis. Also, percent composition of every transcript compo nent of each unigene was calculated.
From these success, the original assembly file made by Trinity was filtered to remove transcripts VX-770 CFTR inhibitor that represent less than 5% on the RSEM primarily based expression amount of its parent unigene or tran scripts with transcripts per million worth beneath 0. five. Coding sequence was predicted through the filtered tran scripts using the transcripts to greatest scoring ORFs. pl script distributed together with the Trinity application from the two strands from the transcripts. This approach makes use of the soft ware Transdecoder which initially identifies the longest open studying frame for every transcript then employs the 500 longest ORFs to develop a Markov model against a randomization of those ORFs to distinguish amongst coding and non coding areas. This model is then utilised to score the likelihood from the longest ORFs in all the transcripts, reporting only people putative ORFs which outscore the other reading frames. Hence, the minimal abundance filtered transcript assem bly was split into contigs that incorporate full open read through ing frames, contigs containing transcript fragments with predicted partial open reading frames, and contigs con taining no ORF prediction.
